Minor and Major: Penn Study Finds A New Gene Expression Mechanism
A rare, small RNA turns a gene-splicing machine into a switch that controls the expression of hundreds of human genes. Howard Hughes Medical Institute Investigator and professor of Biochemistry Gideon Dreyfuss, PhD, and his team from the Perelman School of Medicine at the University of Pennsylvania, discovered an entirely new aspect of the gene-splicing process that produces messenger RNA (mRNA).
The investigators found that a scarce, small RNA, called U6atac, controls the expression of hundreds of genes that have critical functions in cell growth, cell-cycle control, and global control of physiology. Their results were published in the journal eLife.
These genes encode proteins that play essential roles in cell physiology such as several transcription regulators, ion channels, signaling proteins, and DNA damage-repair proteins. Their levels in cells are regulated by the activity of the splicing machinery, which acts as a valve to control essential regulators of cell growth and response to external stimuli.
Dreyfuss, who studies RNA-binding proteins and their role in such diseases as spinal muscular atrophy and other motor neuron degenerative diseases, describes the findings as “completely unanticipated.”
Complicated Splicing
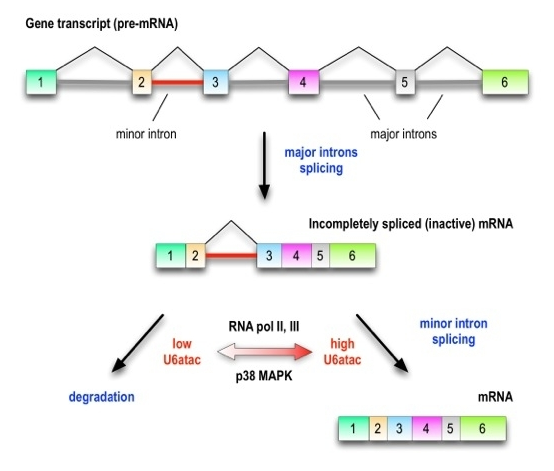
As DNA is transcribed into RNA and then into the various proteins that perform the functions of life, non-coding gene sequences (introns) need to be removed from the transcribed RNA strand and the remaining gene sequences (exons) joined together. This is the job of specialized molecular machinery called the spliceosome. There are two varieties of spliceosomes, the so-called major and minor. The major spliceosome is by far the most abundant, such that the role of its minor counterpart is often disregarded.
“Most of the time the minor spliceosome, which has similar but not identical components to that of the major, isn’t even mentioned,” says Dreyfuss. With each type of spliceosome recognizing different splicing cues, the major spliceosome acts on the vast majority of introns (>200,000) and the minor one splices the several hundred minor-type introns.
But the evolutionary persistence and role of the minor spliceosome has been a puzzle to scientists, since the minor introns it targets are far outnumbered by the major introns handled by the major spliceosome, and the minor spliceosome is often inefficient. But the mRNAs produced from genes that have a minor intron are not ready until all their introns, both major and minor, are spliced. Thus a single inefficiently spliced minor intron can hold up expression – mRNA and protein production – for an entire gene. Researchers have therefore wondered why the apparently superfluous minor spliceosome hasn’t been eliminated altogether through normal evolution.
“One looks at it and asks, we’ve known that minor spliceosomes are inefficient, why even bother to keep them under evolution’s relentless selection pressure?” notes Dreyfuss. “It’s been difficult to rationalize the conservation of minor introns and the minor spliceosome on the basis of splicing alone, as with few cue changes this function could simply have been performed by the major spliceosome.”
Click here to view the full release.